GRK2 Antibody - #DF6538
| Product: | GRK2 Antibody |
| Catalog: | DF6538 |
| Description: | Rabbit polyclonal antibody to GRK2 |
| Application: | WB IHC IF/ICC |
| Cited expt.: | WB |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Pig, Bovine, Horse, Sheep, Dog, Chicken |
| Mol.Wt.: | 80kDa; 80kD(Calculated). |
| Uniprot: | P25098 |
| RRID: | AB_2838500 |
Related Downloads
Protocols
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# DF6538, RRID:AB_2838500.
Fold/Unfold
ADRBK1; Adrenergic beta receptor kinase 1; ARBK1_HUMAN; BARK; BARK1; Beta adrenergic receptor kinase 1; Beta ARK 1; Beta ARK1; Beta-adrenergic receptor kinase 1; Beta-ARK-1; FLJ16718; G protein coupled receptor kinase 2; G-protein coupled receptor kinase 2; GRK2;
Immunogens
A synthesized peptide derived from human GRK2, corresponding to a region within the internal amino acids.
- P25098 ARBK1_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MADLEAVLADVSYLMAMEKSKATPAARASKKILLPEPSIRSVMQKYLEDRGEVTFEKIFSQKLGYLLFRDFCLNHLEEARPLVEFYEEIKKYEKLETEEERVARSREIFDSYIMKELLACSHPFSKSATEHVQGHLGKKQVPPDLFQPYIEEICQNLRGDVFQKFIESDKFTRFCQWKNVELNIHLTMNDFSVHRIIGRGGFGEVYGCRKADTGKMYAMKCLDKKRIKMKQGETLALNERIMLSLVSTGDCPFIVCMSYAFHTPDKLSFILDLMNGGDLHYHLSQHGVFSEADMRFYAAEIILGLEHMHNRFVVYRDLKPANILLDEHGHVRISDLGLACDFSKKKPHASVGTHGYMAPEVLQKGVAYDSSADWFSLGCMLFKLLRGHSPFRQHKTKDKHEIDRMTLTMAVELPDSFSPELRSLLEGLLQRDVNRRLGCLGRGAQEVKESPFFRSLDWQMVFLQKYPPPLIPPRGEVNAADAFDIGSFDEEDTKGIKLLDSDQELYRNFPLTISERWQQEVAETVFDTINAETDRLEARKKAKNKQLGHEEDYALGKDCIMHGYMSKMGNPFLTQWQRRYFYLFPNRLEWRGEGEAPQSLLTMEEIQSVEETQIKERKCLLLKIRGGKQFILQCDSDPELVQWKKELRDAYREAQQLVQRVPKMKNKPRSPVVELSKVPLVQRGSANGL
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Specifically phosphorylates the agonist-occupied form of the beta-adrenergic and closely related receptors, probably inducing a desensitization of them. Key regulator of LPAR1 signaling. Competes with RALA for binding to LPAR1 thus affecting the signaling properties of the receptor. Desensitizes LPAR1 and LPAR2 in a phosphorylation-independent manner. Positively regulates ciliary smoothened (SMO)-dependent Hedgehog (Hh) signaling pathway by facilitating the trafficking of SMO into the cilium and the stimulation of SMO activity (By similarity).
Cytoplasm. Cell membrane.
Expressed in peripheral blood leukocytes.
The PH domain binds anionic phospholipids and helps recruiting ADRBK1 from the cytoplasm to plasma membrane close to activated receptors. It mediates binding to G protein beta and gamma subunits, competing with G-alpha subunits and other G-betagamma effectors.
Belongs to the protein kinase superfamily. AGC Ser/Thr protein kinase family. GPRK subfamily.
Research Fields
· Cellular Processes > Transport and catabolism > Endocytosis. (View pathway)
· Environmental Information Processing > Signal transduction > Hedgehog signaling pathway. (View pathway)
· Human Diseases > Substance dependence > Morphine addiction.
· Organismal Systems > Immune system > Chemokine signaling pathway. (View pathway)
· Organismal Systems > Nervous system > Glutamatergic synapse.
· Organismal Systems > Sensory system > Olfactory transduction.
References
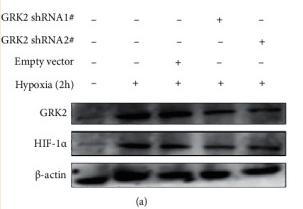
Application: WB Species: rat Sample: Bone marrow–derived macrophages
Application: WB Species: Human Sample:
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.