Occludin Antibody - #DF7504
| Product: | Occludin Antibody |
| Catalog: | DF7504 |
| Description: | Rabbit polyclonal antibody to Occludin |
| Application: | WB IHC IF/ICC |
| Cited expt.: | WB, IHC, IF/ICC |
| Reactivity: | Human, Mouse, Rat, Pig |
| Prediction: | Pig, Bovine, Horse, Sheep, Rabbit, Dog, Chicken |
| Mol.Wt.: | 23~30kD, 52~65kD; 59kD(Calculated). |
| Uniprot: | Q16625 |
| RRID: | AB_2841004 |
Related Downloads
Protocols
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# DF7504, RRID:AB_2841004.
Fold/Unfold
BLCPMG; FLJ08163; FLJ18079; FLJ77961; FLJ94056; MGC34277; Occludin; Ocln; OCLN_HUMAN; Tight junction protein occludin;
Immunogens
A synthesized peptide derived from human Occludin, corresponding to a region within the internal amino acids.
Localized at tight junctions of both epithelial and endothelial cells. Highly expressed in kidney. Not detected in testis.
- Q16625 OCLN_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MSSRPLESPPPYRPDEFKPNHYAPSNDIYGGEMHVRPMLSQPAYSFYPEDEILHFYKWTSPPGVIRILSMLIIVMCIAIFACVASTLAWDRGYGTSLLGGSVGYPYGGSGFGSYGSGYGYGYGYGYGYGGYTDPRAAKGFMLAMAAFCFIAALVIFVTSVIRSEMSRTRRYYLSVIIVSAILGIMVFIATIVYIMGVNPTAQSSGSLYGSQIYALCNQFYTPAATGLYVDQYLYHYCVVDPQEAIAIVLGFMIIVAFALIIFFAVKTRRKMDRYDKSNILWDKEHIYDEQPPNVEEWVKNVSAGTQDVPSPPSDYVERVDSPMAYSSNGKVNDKRFYPESSYKSTPVPEVVQELPLTSPVDDFRQPRYSSGGNFETPSKRAPAKGRAGRSKRTEQDHYETDYTTGGESCDELEEDWIREYPPITSDQQRQLYKRNFDTGLQEYKSLQSELDEINKELSRLDKELDDYREESEEYMAAADEYNRLKQVKGSADYKSKKNHCKQLKSKLSHIKKMVGDYDRQKT
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
May play a role in the formation and regulation of the tight junction (TJ) paracellular permeability barrier. It is able to induce adhesion when expressed in cells lacking tight junctions.
Dephosphorylated by PTPRJ. The tyrosine phosphorylation on Tyr-398 and Tyr-402 reduces its ability to interact with TJP1. Phosphorylation at Ser-490 also attenuates the interaction with TJP1.
Cell membrane>Multi-pass membrane protein. Cell junction>Tight junction.
Localized at tight junctions of both epithelial and endothelial cells. Highly expressed in kidney. Not detected in testis.
The C-terminal is cytoplasmic and is important for interaction with ZO-1. Sufficient for the tight junction localization. Involved in the regulation of the permeability barrier function of the tight junction (By similarity). The first extracellular loop participates in an adhesive interaction.
Belongs to the ELL/occludin family.
Research Fields
· Cellular Processes > Cellular community - eukaryotes > Tight junction. (View pathway)
· Environmental Information Processing > Signaling molecules and interaction > Cell adhesion molecules (CAMs). (View pathway)
· Human Diseases > Infectious diseases: Bacterial > Pathogenic Escherichia coli infection.
· Human Diseases > Infectious diseases: Viral > Hepatitis C.
· Organismal Systems > Immune system > Leukocyte transendothelial migration. (View pathway)
References
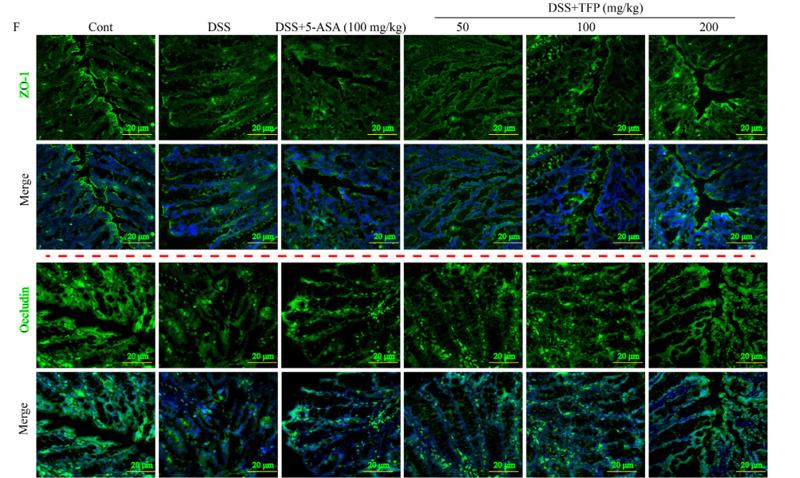
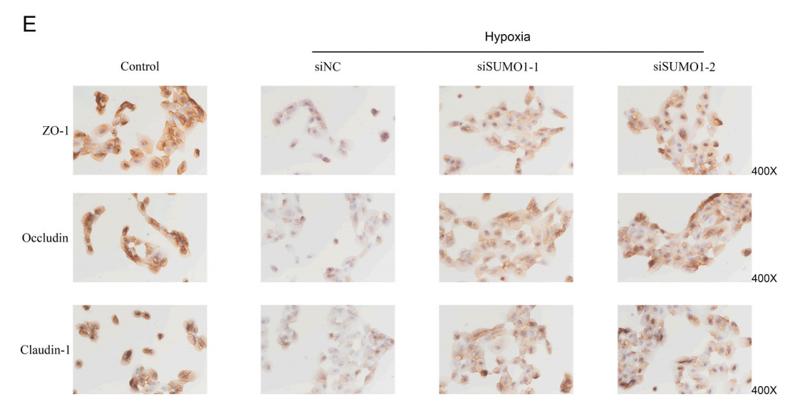
Application: IF/ICC Species: Mouse Sample:
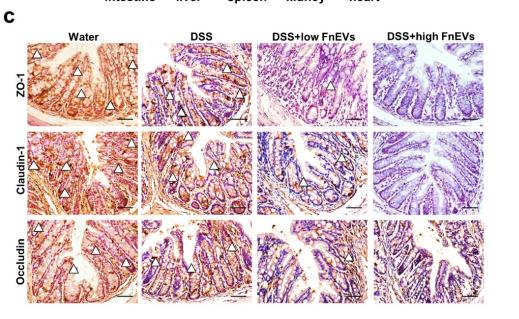
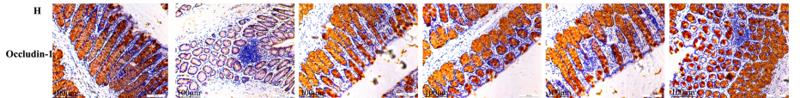
Application: IHC Species: mouse Sample: colon
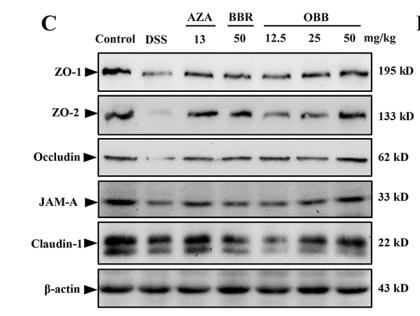
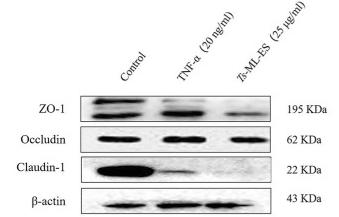
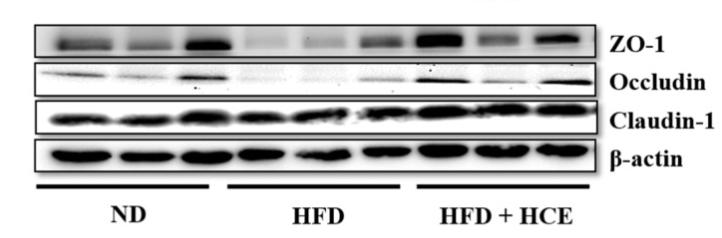
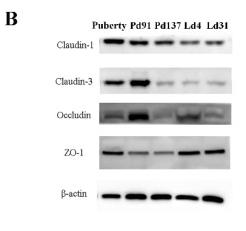
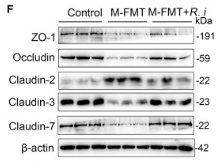
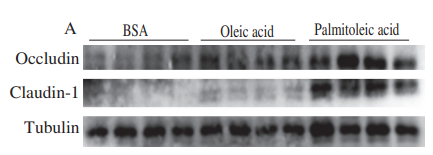
Application: WB Species: Mouse Sample:
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.