Cytokeratin 7 Antibody - #AF0195
| Product: | Cytokeratin 7 Antibody |
| Catalog: | AF0195 |
| Description: | Rabbit polyclonal antibody to Cytokeratin 7 |
| Application: | WB IHC IF/ICC |
| Cited expt.: | WB, IHC, IF/ICC |
| Reactivity: | Human |
| Mol.Wt.: | 51kDa; 51kD(Calculated). |
| Uniprot: | P08729 |
| RRID: | AB_2833388 |
Related Downloads
Protocols
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF0195, RRID:AB_2833388.
Fold/Unfold
CK 7; CK-7; CK7; Cytokeratin 7; Cytokeratin-7; D15Wsu77e; K2C7; K2C7_HUMAN; K7; Keratin 7; Keratin 7, type II; Keratin type II cytoskeletal 7; Keratin, 55K type II cytoskeletal; Keratin, simple epithelial; Keratin, simple epithelial type I, K7; Keratin, type II cytoskeletal 7; Keratin-7; Krt2-7; KRT7; MGC11625; MGC129731; MGC3625; Sarcolectin; SCL; Type II mesothelial keratin K7; Type-II keratin Kb7;
Immunogens
A synthesized peptide derived from human Cytokeratin 7, corresponding to a region within C-terminal amino acids.
Expressed in cultured epidermal, bronchial and mesothelial cells but absent in colon, ectocervix and liver. Observed throughout the glandular cells in the junction between stomach and esophagus but is absent in the esophagus.
- P08729 K2C7_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MSIHFSSPVFTSRSAAFSGRGAQVRLSSARPGGLGSSSLYGLGASRPRVAVRSAYGGPVGAGIREVTINQSLLAPLRLDADPSLQRVRQEESEQIKTLNNKFASFIDKVRFLEQQNKLLETKWTLLQEQKSAKSSRLPDIFEAQIAGLRGQLEALQVDGGRLEAELRSMQDVVEDFKNKYEDEINHRTAAENEFVVLKKDVDAAYMSKVELEAKVDALNDEINFLRTLNETELTELQSQISDTSVVLSMDNSRSLDLDGIIAEVKAQYEEMAKCSRAEAEAWYQTKFETLQAQAGKHGDDLRNTRNEISEMNRAIQRLQAEIDNIKNQRAKLEAAIAEAEERGELALKDARAKQEELEAALQRGKQDMARQLREYQELMSVKLALDIEIATYRKLLEGEESRLAGDGVGAVNISVMNSTGGSSSGGGIGLTLGGTMGSNALSFSSSAGPGLLKAYSIRTASASRRSARD
Research Backgrounds
Blocks interferon-dependent interphase and stimulates DNA synthesis in cells. Involved in the translational regulation of the human papillomavirus type 16 E7 mRNA (HPV16 E7).
Arg-20 is dimethylated, probably to asymmetric dimethylarginine.
Cytoplasm.
Expressed in cultured epidermal, bronchial and mesothelial cells but absent in colon, ectocervix and liver. Observed throughout the glandular cells in the junction between stomach and esophagus but is absent in the esophagus.
Belongs to the intermediate filament family.
References
Application: WB Species: Human Sample: ovarian cell
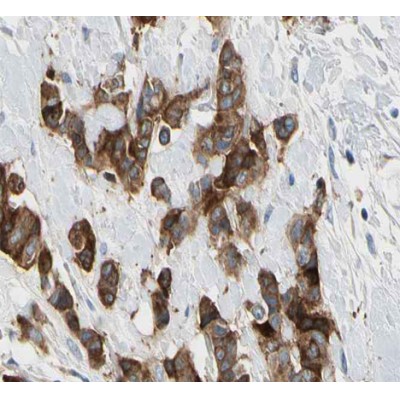
Application: IHC Species: Human Sample: ovarian tumor tissues
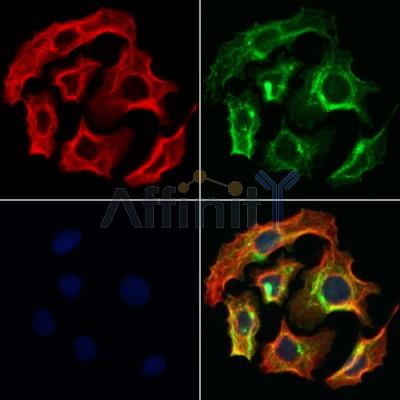
Application: IF/ICC Species: Human Sample:
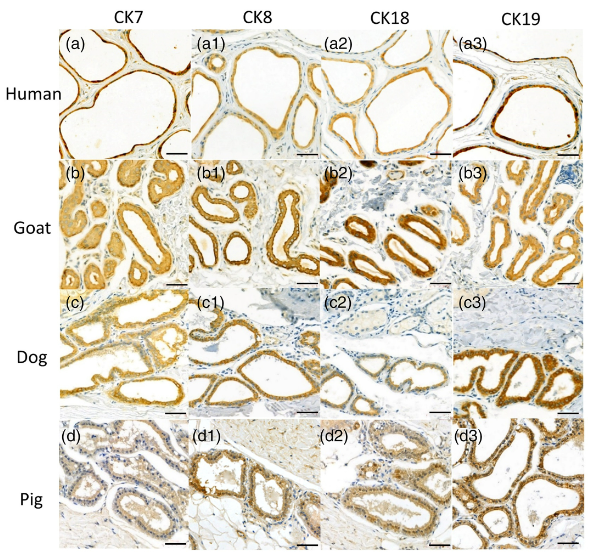
Application: IHC Species: Human,Goat,Dog,Pig Sample: EAC glandular cells
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.