Chk1 Antibody - #AF6007
| Product: | Chk1 Antibody |
| Catalog: | AF6007 |
| Description: | Rabbit polyclonal antibody to Chk1 |
| Application: | WB IHC IF/ICC |
| Cited expt.: | WB |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Pig, Zebrafish, Bovine, Horse, Sheep, Rabbit, Dog, Chicken |
| Mol.Wt.: | 56kDa; 54kD(Calculated). |
| Uniprot: | O14757 |
| RRID: | AB_2834941 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF6007, RRID:AB_2834941.
Fold/Unfold
C85740; Cell cycle checkpoint kinase; Checkpoint , S. pombe, homolog of, 1; Checkpoint kinase 1; Checkpoint kinase 1 homolog (S. pombe); CHEK 1; Chek1; Chk 1; Chk1; CHK1 checkpoint homolog (S. pombe); CHK1_HUMAN; EC 2.7.11.1; rad27; Serine/threonine protein kinase Chk1; Serine/threonine-protein kinase CHK1; STT3, subunit of the oligosaccharyltransferase complex, homolog A (S. cerevisiae);
Immunogens
A synthesized peptide derived from human Chk1, corresponding to a region within the internal amino acids.
Expressed ubiquitously with the most abundant expression in thymus, testis, small intestine and colon.
- O14757 CHK1_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MAVPFVEDWDLVQTLGEGAYGEVQLAVNRVTEEAVAVKIVDMKRAVDCPENIKKEICINKMLNHENVVKFYGHRREGNIQYLFLEYCSGGELFDRIEPDIGMPEPDAQRFFHQLMAGVVYLHGIGITHRDIKPENLLLDERDNLKISDFGLATVFRYNNRERLLNKMCGTLPYVAPELLKRREFHAEPVDVWSCGIVLTAMLAGELPWDQPSDSCQEYSDWKEKKTYLNPWKKIDSAPLALLHKILVENPSARITIPDIKKDRWYNKPLKKGAKRPRVTSGGVSESPSGFSKHIQSNLDFSPVNSASSEENVKYSSSQPEPRTGLSLWDTSPSYIDKLVQGISFSQPTCPDHMLLNSQLLGTPGSSQNPWQRLVKRMTRFFTKLDADKSYQCLKETCEKLGYQWKKSCMNQVTISTTDRRNNKLIFKVNLLEMDDKILVDFRLSKGDGLEFKRHFLKIKGKLIDIVSSQKIWLPAT
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Serine/threonine-protein kinase which is required for checkpoint-mediated cell cycle arrest and activation of DNA repair in response to the presence of DNA damage or unreplicated DNA. May also negatively regulate cell cycle progression during unperturbed cell cycles. This regulation is achieved by a number of mechanisms that together help to preserve the integrity of the genome. Recognizes the substrate consensus sequence [R-X-X-S/T]. Binds to and phosphorylates CDC25A, CDC25B and CDC25C. Phosphorylation of CDC25A at 'Ser-178' and 'Thr-507' and phosphorylation of CDC25C at 'Ser-216' creates binding sites for 14-3-3 proteins which inhibit CDC25A and CDC25C. Phosphorylation of CDC25A at 'Ser-76', 'Ser-124', 'Ser-178', 'Ser-279' and 'Ser-293' promotes proteolysis of CDC25A. Phosphorylation of CDC25A at 'Ser-76' primes the protein for subsequent phosphorylation at 'Ser-79', 'Ser-82' and 'Ser-88' by NEK11, which is required for polyubiquitination and degradation of CDCD25A. Inhibition of CDC25 leads to increased inhibitory tyrosine phosphorylation of CDK-cyclin complexes and blocks cell cycle progression. Also phosphorylates NEK6. Binds to and phosphorylates RAD51 at 'Thr-309', which promotes the release of RAD51 from BRCA2 and enhances the association of RAD51 with chromatin, thereby promoting DNA repair by homologous recombination. Phosphorylates multiple sites within the C-terminus of TP53, which promotes activation of TP53 by acetylation and promotes cell cycle arrest and suppression of cellular proliferation. Also promotes repair of DNA cross-links through phosphorylation of FANCE. Binds to and phosphorylates TLK1 at 'Ser-743', which prevents the TLK1-dependent phosphorylation of the chromatin assembly factor ASF1A. This may enhance chromatin assembly both in the presence or absence of DNA damage. May also play a role in replication fork maintenance through regulation of PCNA. May regulate the transcription of genes that regulate cell-cycle progression through the phosphorylation of histones. Phosphorylates histone H3.1 (to form H3T11ph), which leads to epigenetic inhibition of a subset of genes. May also phosphorylate RB1 to promote its interaction with the E2F family of transcription factors and subsequent cell cycle arrest.
Endogenous repressor of isoform 1, interacts with, and antagonizes CHK1 to promote the S to G2/M phase transition.
Phosphorylated by ATR in a RAD17-dependent manner in response to ultraviolet irradiation and inhibition of DNA replication. Phosphorylated by ATM in response to ionizing irradiation. ATM and ATR can both phosphorylate Ser-317 and Ser-345 and this results in enhanced kinase activity. Phosphorylation at Ser-345 induces a change in the conformation of the protein, activates the kinase activity and is a prerequisite for interaction with FBXO6 and subsequent ubiquitination at Lys-436. Phosphorylation at Ser-345 also increases binding to 14-3-3 proteins and promotes nuclear retention. Conversely, dephosphorylation at Ser-345 by PPM1D may contribute to exit from checkpoint mediated cell cycle arrest. Phosphorylation at Ser-280 by AKT1/PKB, may promote mono and/or diubiquitination. Also phosphorylated at undefined residues during mitotic arrest, resulting in decreased activity.
Ubiquitinated. Mono or diubiquitination promotes nuclear exclusion (By similarity). The activated form (phosphorylated on Ser-345) is polyubiquitinated at Lys-436 by some SCF-type E3 ubiquitin ligase complex containing FBXO6 promoting its degradation. Ubiquitination and degradation are required to terminate the checkpoint and ensure that activated CHEK1 does not accumulate as cells progress through S phase, when replication forks encounter transient impediments during normal DNA replication.
Nucleus. Cytoplasm. Cytoplasm>Cytoskeleton>Microtubule organizing center>Centrosome.
Note: Nuclear export is mediated at least in part by XPO1/CRM1. Also localizes to the centrosome specifically during interphase, where it may protect centrosomal CDC2 kinase from inappropriate activation by cytoplasmic CDC25B.
Expressed ubiquitously with the most abundant expression in thymus, testis, small intestine and colon.
The autoinhibitory region (AIR) inhibits the activity of the kinase domain.
Belongs to the protein kinase superfamily. CAMK Ser/Thr protein kinase family. NIM1 subfamily.
Research Fields
· Cellular Processes > Cell growth and death > Cell cycle. (View pathway)
· Cellular Processes > Cell growth and death > p53 signaling pathway. (View pathway)
· Cellular Processes > Cell growth and death > Cellular senescence. (View pathway)
· Human Diseases > Infectious diseases: Viral > HTLV-I infection.
· Human Diseases > Cancers: Overview > Viral carcinogenesis.
References
Application: WB Species: human Sample: DLBCL cells
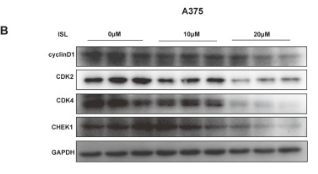
Application: WB Species: Human Sample: A375 and SK-MEL-28 cell
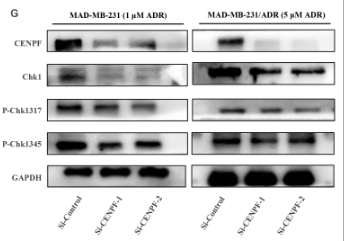
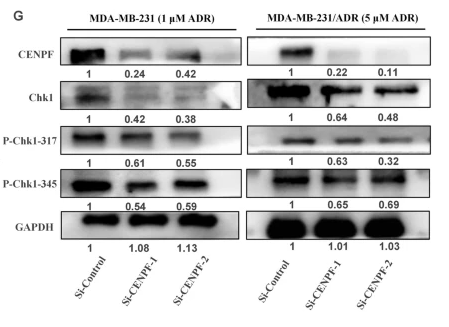
Application: WB Species: Human Sample: MDA-MB-231 and MDA-MB-231/ADR cells
Application: WB Species: Human Sample:
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.