TMS1/ASC Antibody - #DF6304
| Product: | TMS1/ASC Antibody |
| Catalog: | DF6304 |
| Description: | Rabbit polyclonal antibody to TMS1/ASC |
| Application: | WB IHC IF/ICC |
| Cited expt.: | WB, IHC, IF/ICC |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Bovine, Horse, Sheep, Rabbit |
| Mol.Wt.: | 15~25kDa; 22kD(Calculated). |
| Uniprot: | Q9ULZ3 |
| RRID: | AB_2838270 |
Related Downloads
Protocols
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# DF6304, RRID:AB_2838270.
Fold/Unfold
Apoptosis associated speck like protein containing a CARD; Apoptosis-associated speck-like protein containing a CARD; ASC; ASC_HUMAN; CARD 5; CARD5; Caspase recruitment domain containing protein 5; Caspase recruitment domain protein 5; Caspase recruitment domain-containing protein 5; hASC; MGC10332; PYCARD; PYD and CARD domain containing; PYD and CARD domain containing protein; PYD and CARD domain-containing protein; Target of methylation induced silencing 1; Target of methylation-induced silencing 1; TMS 1; TMS; TMS1;
Immunogens
A synthesized peptide derived from human TMS1/ASC, corresponding to a region within N-terminal amino acids.
Widely expressed at low levels. Detected in peripheral blood leukocytes, lung, small intestine, spleen, thymus, colon and at lower levels in placenta, liver and kidney. Very low expression in skeletal muscle, heart and brain. Expressed in lung epithelial cells (at protein level) (PubMed:23229815). Detected in the leukemia cell lines HL-60 and U-937, but not in Jurkat T-cell lymphoma and Daudi Burkitt's lymphoma. Detected in the melanoma cell line WM35, but not in WM793. Not detected in HeLa cervical carcinoma cells and MOLT-4 lymphocytic leukemia cells.
- Q9ULZ3 ASC_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MGRARDAILDALENLTAEELKKFKLKLLSVPLREGYGRIPRGALLSMDALDLTDKLVSFYLETYGAELTANVLRDMGLQEMAGQLQAATHQGSGAAPAGIQAPPQSAAKPGLHFIDQHRAALIARVTNVEWLLDALYGKVLTDEQYQAVRAEPTNPSKMRKLFSFTPAWNWTCKDLLLQALRESQSYLVEDLERS
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Functions as key mediator in apoptosis and inflammation. Promotes caspase-mediated apoptosis involving predominantly caspase-8 and also caspase-9 in a probable cell type-specific manner. Involved in activation of the mitochondrial apoptotic pathway, promotes caspase-8-dependent proteolytic maturation of BID independently of FADD in certain cell types and also mediates mitochondrial translocation of BAX and activates BAX-dependent apoptosis coupled to activation of caspase-9, -2 and -3. Involved in macrophage pyroptosis, a caspase-1-dependent inflammatory form of cell death and is the major constituent of the ASC pyroptosome which forms upon potassium depletion and rapidly recruits and activates caspase-1. In innate immune response believed to act as an integral adapter in the assembly of the inflammasome which activates caspase-1 leading to processing and secretion of proinflammatory cytokines. The function as activating adapter in different types of inflammasomes is mediated by the pyrin and CARD domains and their homotypic interactions. Required for recruitment of caspase-1 to inflammasomes containing certain pattern recognition receptors, such as NLRP2, NLRP3, AIM2 and probably IFI16. In the NLRP1 and NLRC4 inflammasomes seems not be required but facilitates the processing of procaspase-1. In cooperation with NOD2 involved in an inflammasome activated by bacterial muramyl dipeptide leading to caspase-1 activation. May be involved in DDX58-triggered proinflammatory responses and inflammasome activation. Isoform 2 may have a regulating effect on the function as inflammasome adapter. Isoform 3 seems to inhibit inflammasome-mediated maturation of interleukin-1 beta. In collaboration with AIM2 which detects cytosolic double-stranded DNA may also be involved in a caspase-1-independent cell death that involves caspase-8. In adaptive immunity may be involved in maturation of dendritic cells to stimulate T-cell immunity and in cytoskeletal rearrangements coupled to chemotaxis and antigen uptake may be involved in post-transcriptional regulation of the guanine nucleotide exchange factor DOCK2; the latter function is proposed to involve the nuclear form. Also involved in transcriptional activation of cytokines and chemokines independent of the inflammasome; this function may involve AP-1, NF-kappa-B, MAPK and caspase-8 signaling pathways. For regulation of NF-kappa-B activating and inhibiting functions have been reported. Modulates NF-kappa-B induction at the level of the IKK complex by inhibiting kinase activity of CHUK and IKBK. Proposed to compete with RIPK2 for association with CASP1 thereby down-regulating CASP1-mediated RIPK2-dependent NF-kappa-B activation and activating interleukin-1 beta processing. Modulates host resistance to DNA virus infection, probably by inducing the cleavage of and inactivating CGAS in presence of cytoplasmic double-stranded DNA.
Phosphorylated.
Cytoplasm. Endoplasmic reticulum. Mitochondrion. Nucleus.
Note: Upstream of caspase activation, a redistribution from the cytoplasm to the aggregates occurs. These appear as hollow, perinuclear spherical, ball-like structures. Upon NLRP3 inflammasome activation redistributes to the perinuclear space localizing to endoplasmic reticulum and mitochondria. Localized primarily to the nucleus in resting monocytes/macrophages and rapidly redistributed to the cytoplasm upon pathogen infection. Localized to large cytoplasmic aggregate appearing as a speck containing AIM2, PYCARD, CASP8 and bacterial DNA after infection with Francisella tularensis (By similarity).
Golgi apparatus membrane.
Note: (Microbial infection) Upon HRSV infection, the protein is mainly located in lipid rafts in the Golgi membrane.
Widely expressed at low levels. Detected in peripheral blood leukocytes, lung, small intestine, spleen, thymus, colon and at lower levels in placenta, liver and kidney. Very low expression in skeletal muscle, heart and brain. Expressed in lung epithelial cells (at protein level). Detected in the leukemia cell lines HL-60 and U-937, but not in Jurkat T-cell lymphoma and Daudi Burkitt's lymphoma. Detected in the melanoma cell line WM35, but not in WM793. Not detected in HeLa cervical carcinoma cells and MOLT-4 lymphocytic leukemia cells.
The CARD domain mediates interaction with CASP1 and NLRC4 (PubMed:14634131 and PubMed:11967258).
The pyrin domain mediates homotypic interactions with pyrin domains of proteins such as of NLRP3, PYDC1, PYDC2 and AIM2.
Research Fields
· Cellular Processes > Cell growth and death > Necroptosis. (View pathway)
· Human Diseases > Infectious diseases: Bacterial > Salmonella infection.
· Human Diseases > Infectious diseases: Bacterial > Pertussis.
· Human Diseases > Infectious diseases: Bacterial > Legionellosis.
· Human Diseases > Infectious diseases: Viral > Influenza A.
· Organismal Systems > Immune system > NOD-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > Cytosolic DNA-sensing pathway. (View pathway)
References
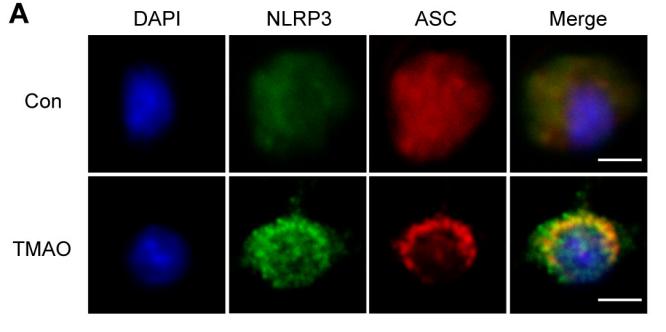
Application: IF/ICC Species: mouse Sample: BMDMs
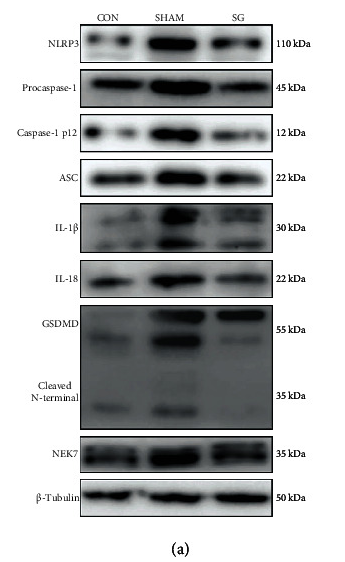
Application: WB Species: Rat Sample:
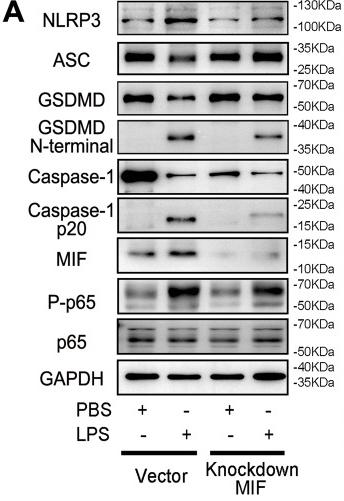
Application: WB Species: Mouse Sample:
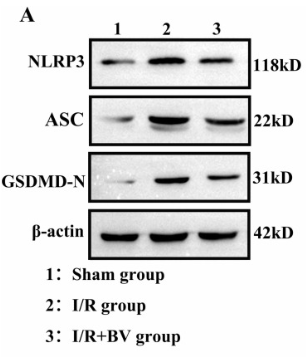
Application: WB Species: Human Sample: HK-2 cells
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.