CD44 Antibody - #AF6186
| Product: | CD44 Antibody |
| Catalog: | AF6186 |
| Description: | Rabbit polyclonal antibody to CD44 |
| Application: | WB IF/ICC |
| Cited expt.: | IF/ICC |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Pig, Bovine, Horse, Sheep, Rabbit, Dog, Chicken, Xenopus |
| Mol.Wt.: | 90kDa; 82kD(Calculated). |
| Uniprot: | P16070 |
| RRID: | AB_2835069 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF6186, RRID:AB_2835069.
Fold/Unfold
LHR; BA-1; CD 44; CD44; CD44 antigen; CD44 molecule (Indian blood group); CD44 molecule; CD44_HUMAN; CDw44; Cell surface glycoprotein CD44; chondroitin sulfate proteoglycan 8; CSPG8; ECMR-III; Epican; Extracellular matrix receptor III; GP90 lymphocyte homing/adhesion receptor; HCELL; hematopoietic cell E- and L-selectin ligand; Heparan sulfate proteoglycan; Hermes antigen; homing function and Indian blood group system; HSA; HUTCH-I; HUTCH1; Hyaluronate receptor; IN; INLU-related p80 Glycoprotein; MC56; MDU2; MDU3; MGC10468; MIC4; MUTCH1; PGP-1; PGP-I; PGP1; Phagocytic glycoprotein 1; Phagocytic glycoprotein I; Soluble CD44;
Immunogens
A synthesized peptide derived from human CD44, corresponding to a region within C-terminal amino acids.
Isoform 10 (epithelial isoform) is expressed by cells of epithelium and highly expressed by carcinomas. Expression is repressed in neuroblastoma cells.
- P16070 CD44_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MDKFWWHAAWGLCLVPLSLAQIDLNITCRFAGVFHVEKNGRYSISRTEAADLCKAFNSTLPTMAQMEKALSIGFETCRYGFIEGHVVIPRIHPNSICAANNTGVYILTSNTSQYDTYCFNASAPPEEDCTSVTDLPNAFDGPITITIVNRDGTRYVQKGEYRTNPEDIYPSNPTDDDVSSGSSSERSSTSGGYIFYTFSTVHPIPDEDSPWITDSTDRIPATTLMSTSATATETATKRQETWDWFSWLFLPSESKNHLHTTTQMAGTSSNTISAGWEPNEENEDERDRHLSFSGSGIDDDEDFISSTISTTPRAFDHTKQNQDWTQWNPSHSNPEVLLQTTTRMTDVDRNGTTAYEGNWNPEAHPPLIHHEHHEEEETPHSTSTIQATPSSTTEETATQKEQWFGNRWHEGYRQTPKEDSHSTTGTAAASAHTSHPMQGRTTPSPEDSSWTDFFNPISHPMGRGHQAGRRMDMDSSHSITLQPTANPNTGLVEDLDRTGPLSMTTQQSNSQSFSTSHEGLEEDKDHPTTSTLTSSNRNDVTGGRRDPNHSEGSTTLLEGYTSHYPHTKESRTFIPVTSAKTGSFGVTAVTVGDSNSNVNRSLSGDQDTFHPSGGSHTTHGSESDGHSHGSQEGGANTTSGPIRTPQIPEWLIILASLLALALILAVCIAVNSRRRCGQKKKLVINSGNGAVEDRKPSGLNGEASKSQEMVHLVNKESSETPDQFMTADETRNLQNVDMKIGV
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Cell-surface receptor that plays a role in cell-cell interactions, cell adhesion and migration, helping them to sense and respond to changes in the tissue microenvironment. Participates thereby in a wide variety of cellular functions including the activation, recirculation and homing of T-lymphocytes, hematopoiesis, inflammation and response to bacterial infection. Engages, through its ectodomain, extracellular matrix components such as hyaluronan/HA, collagen, growth factors, cytokines or proteases and serves as a platform for signal transduction by assembling, via its cytoplasmic domain, protein complexes containing receptor kinases and membrane proteases. Such effectors include PKN2, the RhoGTPases RAC1 and RHOA, Rho-kinases and phospholipase C that coordinate signaling pathways promoting calcium mobilization and actin-mediated cytoskeleton reorganization essential for cell migration and adhesion.
Proteolytically cleaved in the extracellular matrix by specific proteinases (possibly MMPs) in several cell lines and tumors.
N-glycosylated.
O-glycosylated. O-glycosylation contains more-or-less-sulfated chondroitin sulfate glycans, whose number may affect the accessibility of specific proteinases to their cleavage site(s). It is uncertain if O-glycosylation occurs on Thr-637 or Thr-638.
Phosphorylated; activation of PKC results in the dephosphorylation of Ser-706 (constitutive phosphorylation site), and the phosphorylation of Ser-672.
Cell membrane>Single-pass type I membrane protein. Cell projection>Microvillus.
Note: Colocalizes with actin in membrane protrusions at wounding edges. Co-localizes with RDX, EZR and MSN in microvilli. Localizes to cholesterol-rich membrane-bound lipid raft domains.
Isoform 10 (epithelial isoform) is expressed by cells of epithelium and highly expressed by carcinomas. Expression is repressed in neuroblastoma cells.
The lectin-like LINK domain is responsible for hyaluronan binding.
Research Fields
· Environmental Information Processing > Signaling molecules and interaction > ECM-receptor interaction. (View pathway)
· Human Diseases > Infectious diseases: Bacterial > Shigellosis.
· Human Diseases > Infectious diseases: Viral > Epstein-Barr virus infection.
· Human Diseases > Cancers: Overview > Proteoglycans in cancer.
· Human Diseases > Cancers: Overview > MicroRNAs in cancer.
· Organismal Systems > Immune system > Hematopoietic cell lineage. (View pathway)
References
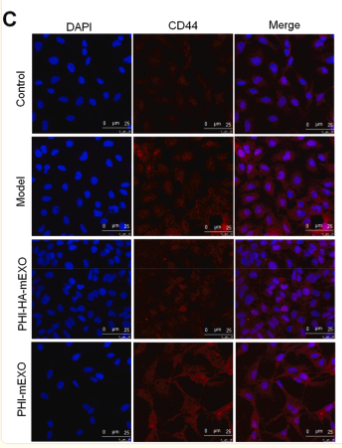
Application: IF/ICC Species: zebrafish Sample: LX2 cells
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.