CCR5 Antibody - #AF6339
| Product: | CCR5 Antibody |
| Catalog: | AF6339 |
| Description: | Rabbit polyclonal antibody to CCR5 |
| Application: | WB IF/ICC |
| Cited expt.: | WB, IF/ICC |
| Reactivity: | Human, Mouse, Rat, Monkey |
| Prediction: | Pig, Bovine, Rabbit, Chicken |
| Mol.Wt.: | 40kDa; 41kD(Calculated). |
| Uniprot: | P51681 |
| RRID: | AB_2835195 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF6339, RRID:AB_2835195.
Fold/Unfold
AM4 7; C C chemokine receptor type 5; C C CKR 5; C-C chemokine receptor type 5; C-C CKR-5; C-C motif chemokine receptor 5 A159A; CC Chemokine Receptor 5; CC Chemokine Receptor Type 5; CC CKR 5; CC-CKR-5; CCCKR 5; CCCKR5; CCR 5; CCR-5; CCR5; CCR5 chemokine (C C motif) receptor 5; CCR5_HUMAN; CD 195; CD195; CD195 Antigen; Chemokine C C motif receptor 5; Chemokine receptor CCR5; CHEMR13; CKR 5; CKR5; CMKBR 5; CMKBR5; FLJ78003; HIV 1 Fusion Coreceptor; HIV-1 fusion coreceptor; HIV1 fusion coreceptor; IDDM22; MIP-1 alpha receptor;
Immunogens
A synthesized peptide derived from human CCR5, corresponding to a region within C-terminal amino acids.
Highly expressed in spleen, thymus, in the myeloid cell line THP-1, in the promyeloblastic cell line KG-1a and on CD4+ and CD8+ T-cells. Medium levels in peripheral blood leukocytes and in small intestine. Low levels in ovary and lung.
- P51681 CCR5_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MDYQVSSPIYDINYYTSEPCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINCKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPGIIFTRSQKEGLHYTCSSHFPYSQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRCRNEKKRHRAVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEKFRNYLLVFFQKHIAKRFCKCCSIFQQEAPERASSVYTRSTGEQEISVGL
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Receptor for a number of inflammatory CC-chemokines including CCL3/MIP-1-alpha, CCL4/MIP-1-beta and RANTES and subsequently transduces a signal by increasing the intracellular calcium ion level. May play a role in the control of granulocytic lineage proliferation or differentiation.
(Microbial infection) Acts as a coreceptor (CD4 being the primary receptor) of human immunodeficiency virus-1/HIV-1.
Sulfated on at least 2 of the N-terminal tyrosines. Sulfation contributes to the efficiency of HIV-1 entry and is required for efficient binding of the chemokines, CCL3 and CCL4.
O-glycosylated, but not N-glycosylated. Ser-6 appears to be the major site. Also sialylated glycans present which contribute to chemokine binding. Thr-16 and Ser-17 may also be glycosylated and, if so, with small moieties such as a T-antigen.
Palmitoylation in the C-terminal is important for cell surface expression, and to a lesser extent, for HIV entry.
Phosphorylation on serine residues in the C-terminal is stimulated by binding CC chemokines especially by APO-RANTES.
Cell membrane>Multi-pass membrane protein.
Highly expressed in spleen, thymus, in the myeloid cell line THP-1, in the promyeloblastic cell line KG-1a and on CD4+ and CD8+ T-cells. Medium levels in peripheral blood leukocytes and in small intestine. Low levels in ovary and lung.
Belongs to the G-protein coupled receptor 1 family.
Research Fields
· Cellular Processes > Transport and catabolism > Endocytosis. (View pathway)
· Environmental Information Processing > Signaling molecules and interaction > Cytokine-cytokine receptor interaction. (View pathway)
· Human Diseases > Infectious diseases: Parasitic > Toxoplasmosis.
· Human Diseases > Cancers: Overview > Viral carcinogenesis.
· Organismal Systems > Immune system > Chemokine signaling pathway. (View pathway)
References
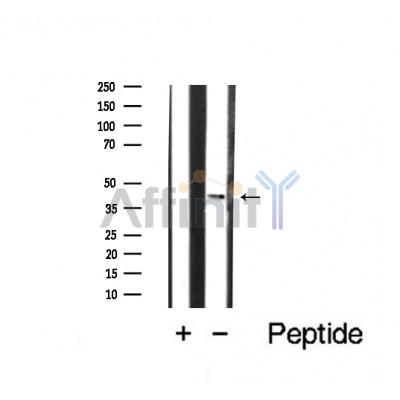
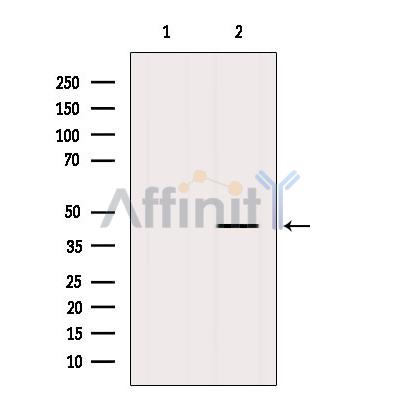
Application: WB Species: human Sample: prostate cancer cells
Application: IF/ICC Species: Mouse Sample:
Application: IHC Species: human Sample: HCC and para-carcinoma live tissues
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.