Phospho-Glycogen Synthase (Ser645) Antibody - #AF3165
| Product: | Phospho-Glycogen Synthase (Ser645) Antibody |
| Catalog: | AF3165 |
| Description: | Rabbit polyclonal antibody to Phospho-Glycogen Synthase (Ser645) |
| Application: | WB IHC IF/ICC |
| Cited expt.: | WB |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Pig, Zebrafish, Bovine, Horse, Sheep, Rabbit, Dog |
| Mol.Wt.: | 83,89kDa; 84kD(Calculated). |
| Uniprot: | P13807 |
| RRID: | AB_2834500 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF3165, RRID:AB_2834500.
Fold/Unfold
Glycogen [starch] synthase; Glycogen synthase 1 (muscle); Glycogen synthase 1; GSY; GYS; Gys1; GYS1_HUMAN; muscle;
Immunogens
A synthesized peptide derived from human Glycogen Synthase around the phosphorylation site of Ser645.
- P13807 GYS1_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MPLNRTLSMSSLPGLEDWEDEFDLENAVLFEVAWEVANKVGGIYTVLQTKAKVTGDEWGDNYFLVGPYTEQGVRTQVELLEAPTPALKRTLDSMNSKGCKVYFGRWLIEGGPLVVLLDVGASAWALERWKGELWDTCNIGVPWYDREANDAVLFGFLTTWFLGEFLAQSEEKPHVVAHFHEWLAGVGLCLCRARRLPVATIFTTHATLLGRYLCAGAVDFYNNLENFNVDKEAGERQIYHRYCMERAAAHCAHVFTTVSQITAIEAQHLLKRKPDIVTPNGLNVKKFSAMHEFQNLHAQSKARIQEFVRGHFYGHLDFNLDKTLYFFIAGRYEFSNKGADVFLEALARLNYLLRVNGSEQTVVAFFIMPARTNNFNVETLKGQAVRKQLWDTANTVKEKFGRKLYESLLVGSLPDMNKMLDKEDFTMMKRAIFATQRQSFPPVCTHNMLDDSSDPILTTIRRIGLFNSSADRVKVIFHPEFLSSTSPLLPVDYEEFVRGCHLGVFPSYYEPWGYTPAECTVMGIPSISTNLSGFGCFMEEHIADPSAYGIYILDRRFRSLDDSCSQLTSFLYSFCQQSRRQRIIQRNRTERLSDLLDWKYLGRYYMSARHMALSKAFPEHFTYEPNEADAAQGYRYPRPASVPPSPSLSRHSSPHQSEDEEDPRNGPLEEDGERYDEDEEAAKDRRNIRAPEWPRRASCTSSTSGSKRNSVDTATSSSLSTPSEPLSPTSSLGEERN
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Transfers the glycosyl residue from UDP-Glc to the non-reducing end of alpha-1,4-glucan.
Phosphorylation at Ser-8 by AMPK inactivates the enzyme activity. Primed phosphorylation at Ser-657 (site 5) by CSNK2A1 and CSNK2A2 is required for inhibitory phosphorylation at Ser-641 (site 3a), Ser-645 (site 3b), Ser-649 (site 3c) and Ser-653 (site 4) by GSK3A an GSK3B (By similarity). Phosphorylated at Ser-641 by DYRK2, leading to inactivation (By similarity). Phosphorylated at Ser-641 by PASK, leading to inactivation; phosphorylation by PASK is inhibited by glycogen. Dephosphorylation at Ser-641 and Ser-645 by PP1 activates the enzyme.
Belongs to the glycosyltransferase 3 family.
Research Fields
· Environmental Information Processing > Signal transduction > PI3K-Akt signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > AMPK signaling pathway. (View pathway)
· Human Diseases > Endocrine and metabolic diseases > Insulin resistance.
· Metabolism > Carbohydrate metabolism > Starch and sucrose metabolism.
· Metabolism > Global and overview maps > Metabolic pathways.
· Organismal Systems > Endocrine system > Insulin signaling pathway. (View pathway)
· Organismal Systems > Endocrine system > Glucagon signaling pathway.
References
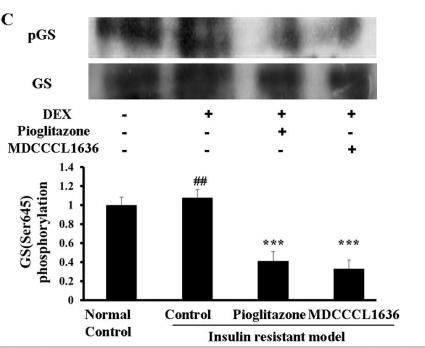
Application: WB Species: human Sample: HepG2 cells
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.