Phospho-PDGF Receptor beta (Tyr740) Antibody - #AF3132
| Product: | Phospho-PDGF Receptor beta (Tyr740) Antibody |
| Catalog: | AF3132 |
| Description: | Rabbit polyclonal antibody to Phospho-PDGF Receptor beta (Tyr740) |
| Application: | WB IHC |
| Cited expt.: | WB |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Pig, Zebrafish, Horse, Sheep, Rabbit, Dog, Chicken, Xenopus |
| Mol.Wt.: | 170kDa; 124kD(Calculated). |
| Uniprot: | P09619 |
| RRID: | AB_2834567 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF3132, RRID:AB_2834567.
Fold/Unfold
Beta platelet derived growth factor receptor; Beta-type platelet-derived growth factor receptor; CD 140B; CD140 antigen-like family member B; CD140b; CD140b antigen; IBGC4; IMF1; JTK12; OTTHUMP00000160528; PDGF R beta; PDGF-R-beta; PDGFR 1; PDGFR; PDGFR beta; PDGFR1; PDGFRB; PGFRB_HUMAN; Platelet derived growth factor receptor 1; Platelet derived growth factor receptor beta; Platelet derived growth factor receptor beta polypeptide;
Immunogens
A synthesized peptide derived from human PDGF Receptor beta around the phosphorylation site of Tyr740.
- P09619 PGFRB_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MRLPGAMPALALKGELLLLSLLLLLEPQISQGLVVTPPGPELVLNVSSTFVLTCSGSAPVVWERMSQEPPQEMAKAQDGTFSSVLTLTNLTGLDTGEYFCTHNDSRGLETDERKRLYIFVPDPTVGFLPNDAEELFIFLTEITEITIPCRVTDPQLVVTLHEKKGDVALPVPYDHQRGFSGIFEDRSYICKTTIGDREVDSDAYYVYRLQVSSINVSVNAVQTVVRQGENITLMCIVIGNEVVNFEWTYPRKESGRLVEPVTDFLLDMPYHIRSILHIPSAELEDSGTYTCNVTESVNDHQDEKAINITVVESGYVRLLGEVGTLQFAELHRSRTLQVVFEAYPPPTVLWFKDNRTLGDSSAGEIALSTRNVSETRYVSELTLVRVKVAEAGHYTMRAFHEDAEVQLSFQLQINVPVRVLELSESHPDSGEQTVRCRGRGMPQPNIIWSACRDLKRCPRELPPTLLGNSSEEESQLETNVTYWEEEQEFEVVSTLRLQHVDRPLSVRCTLRNAVGQDTQEVIVVPHSLPFKVVVISAILALVVLTIISLIILIMLWQKKPRYEIRWKVIESVSSDGHEYIYVDPMQLPYDSTWELPRDQLVLGRTLGSGAFGQVVEATAHGLSHSQATMKVAVKMLKSTARSSEKQALMSELKIMSHLGPHLNVVNLLGACTKGGPIYIITEYCRYGDLVDYLHRNKHTFLQHHSDKRRPPSAELYSNALPVGLPLPSHVSLTGESDGGYMDMSKDESVDYVPMLDMKGDVKYADIESSNYMAPYDNYVPSAPERTCRATLINESPVLSYMDLVGFSYQVANGMEFLASKNCVHRDLAARNVLICEGKLVKICDFGLARDIMRDSNYISKGSTFLPLKWMAPESIFNSLYTTLSDVWSFGILLWEIFTLGGTPYPELPMNEQFYNAIKRGYRMAQPAHASDEIYEIMQKCWEEKFEIRPPFSQLVLLLERLLGEGYKKKYQQVDEEFLRSDHPAILRSQARLPGFHGLRSPLDTSSVLYTAVQPNEGDNDYIIPLPDPKPEVADEGPLEGSPSLASSTLNEVNTSSTISCDSPLEPQDEPEPEPQLELQVEPEPELEQLPDSGCPAPRAEAEDSFL
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Tyrosine-protein kinase that acts as cell-surface receptor for homodimeric PDGFB and PDGFD and for heterodimers formed by PDGFA and PDGFB, and plays an essential role in the regulation of embryonic development, cell proliferation, survival, differentiation, chemotaxis and migration. Plays an essential role in blood vessel development by promoting proliferation, migration and recruitment of pericytes and smooth muscle cells to endothelial cells. Plays a role in the migration of vascular smooth muscle cells and the formation of neointima at vascular injury sites. Required for normal development of the cardiovascular system. Required for normal recruitment of pericytes (mesangial cells) in the kidney glomerulus, and for normal formation of a branched network of capillaries in kidney glomeruli. Promotes rearrangement of the actin cytoskeleton and the formation of membrane ruffles. Binding of its cognate ligands - homodimeric PDGFB, heterodimers formed by PDGFA and PDGFB or homodimeric PDGFD -leads to the activation of several signaling cascades; the response depends on the nature of the bound ligand and is modulated by the formation of heterodimers between PDGFRA and PDGFRB. Phosphorylates PLCG1, PIK3R1, PTPN11, RASA1/GAP, CBL, SHC1 and NCK1. Activation of PLCG1 leads to the production of the cellular signaling molecules diacylglycerol and inositol 1,4,5-trisphosphate, mobilization of cytosolic Ca(2+) and the activation of protein kinase C. Phosphorylation of PIK3R1, the regulatory subunit of phosphatidylinositol 3-kinase, leads to the activation of the AKT1 signaling pathway. Phosphorylation of SHC1, or of the C-terminus of PTPN11, creates a binding site for GRB2, resulting in the activation of HRAS, RAF1 and down-stream MAP kinases, including MAPK1/ERK2 and/or MAPK3/ERK1. Promotes phosphorylation and activation of SRC family kinases. Promotes phosphorylation of PDCD6IP/ALIX and STAM. Receptor signaling is down-regulated by protein phosphatases that dephosphorylate the receptor and its down-stream effectors, and by rapid internalization of the activated receptor.
Autophosphorylated on tyrosine residues upon ligand binding. Autophosphorylation occurs in trans, i.e. one subunit of the dimeric receptor phosphorylates tyrosine residues on the other subunit. Phosphorylation at Tyr-579, and to a lesser degree, at Tyr-581, is important for interaction with SRC family kinases. Phosphorylation at Tyr-740 and Tyr-751 is important for interaction with PIK3R1. Phosphorylation at Tyr-751 is important for interaction with NCK1. Phosphorylation at Tyr-771 and Tyr-857 is important for interaction with RASA1/GAP. Phosphorylation at Tyr-857 is important for efficient phosphorylation of PLCG1 and PTPN11, resulting in increased phosphorylation of AKT1, MAPK1/ERK2 and/or MAPK3/ERK1, PDCD6IP/ALIX and STAM, and in increased cell proliferation. Phosphorylation at Tyr-1009 is important for interaction with PTPN11. Phosphorylation at Tyr-1009 and Tyr-1021 is important for interaction with PLCG1. Phosphorylation at Tyr-1021 is important for interaction with CBL; PLCG1 and CBL compete for the same binding site. Dephosphorylated by PTPRJ at Tyr-751, Tyr-857, Tyr-1009 and Tyr-1021. Dephosphorylated by PTPN2 at Tyr-579 and Tyr-1021.
N-glycosylated.
Ubiquitinated. After autophosphorylation, the receptor is polyubiquitinated, leading to its degradation.
Cell membrane>Single-pass type I membrane protein. Cytoplasmic vesicle. Lysosome lumen.
Note: After ligand binding, the autophosphorylated receptor is ubiquitinated and internalized, leading to its degradation.
Belongs to the protein kinase superfamily. Tyr protein kinase family. CSF-1/PDGF receptor subfamily.
Research Fields
· Cellular Processes > Cellular community - eukaryotes > Focal adhesion. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Gap junction. (View pathway)
· Cellular Processes > Cell motility > Regulation of actin cytoskeleton. (View pathway)
· Environmental Information Processing > Signal transduction > MAPK signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Ras signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Rap1 signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Calcium signaling pathway. (View pathway)
· Environmental Information Processing > Signaling molecules and interaction > Cytokine-cytokine receptor interaction. (View pathway)
· Environmental Information Processing > Signal transduction > Phospholipase D signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > PI3K-Akt signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Jak-STAT signaling pathway. (View pathway)
· Human Diseases > Drug resistance: Antineoplastic > EGFR tyrosine kinase inhibitor resistance.
· Human Diseases > Infectious diseases: Viral > Human papillomavirus infection.
· Human Diseases > Infectious diseases: Viral > HTLV-I infection.
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Overview > MicroRNAs in cancer.
· Human Diseases > Cancers: Specific types > Glioma. (View pathway)
· Human Diseases > Cancers: Specific types > Prostate cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Melanoma. (View pathway)
· Human Diseases > Cancers: Overview > Central carbon metabolism in cancer. (View pathway)
· Human Diseases > Cancers: Overview > Choline metabolism in cancer. (View pathway)
References
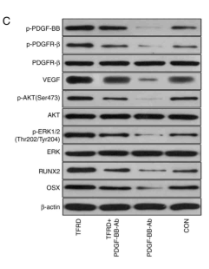
Application: WB Species: Rat Sample:
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.