Phospho-FAK (Tyr397) Antibody - #AF3398
| Product: | Phospho-FAK (Tyr397) Antibody |
| Catalog: | AF3398 |
| Description: | Rabbit polyclonal antibody to Phospho-FAK (Tyr397) |
| Application: | WB IHC IF/ICC |
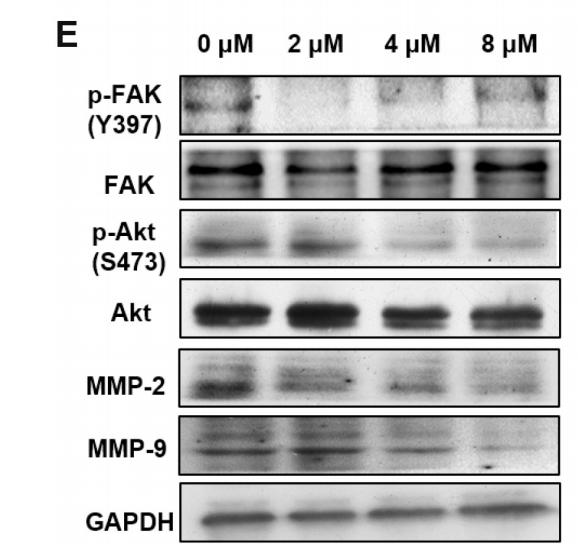
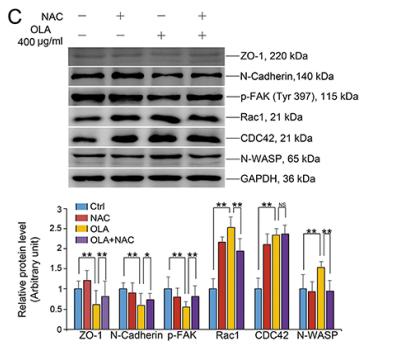
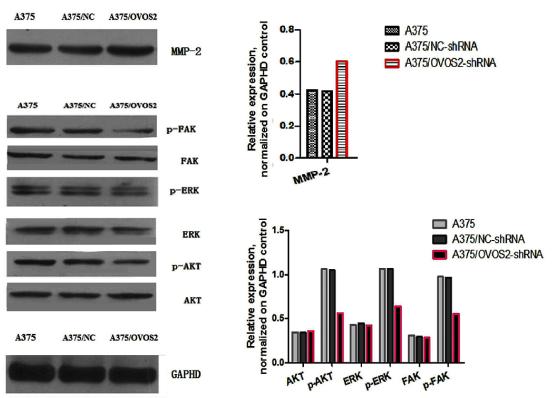
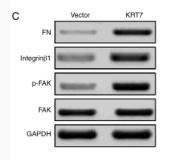
| Cited expt.: | WB, IHC |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Pig, Horse, Sheep, Rabbit, Dog, Chicken, Xenopus |
| Mol.Wt.: | 119kDa; 119kD(Calculated). |
| Uniprot: | Q05397 |
| RRID: | AB_2834829 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF3398, RRID:AB_2834829.
Fold/Unfold
FADK 1; FADK; FAK related non kinase polypeptide; FAK1; FAK1_HUMAN; Focal adhesion kinase 1; Focal adhesion Kinase; Focal adhesion kinase isoform FAK Del33; Focal adhesion kinase related nonkinase; FRNK; p125FAK; pp125FAK; PPP1R71; Protein phosphatase 1 regulatory subunit 71; Protein tyrosine kinase 2; Protein-tyrosine kinase 2; Ptk2; PTK2 protein tyrosine kinase 2;
Immunogens
A synthesized peptide derived from human FAK around the phosphorylation site of Tyr397.
Detected in B and T-lymphocytes. Isoform 1 and isoform 6 are detected in lung fibroblasts (at protein level). Ubiquitous.
- Q05397 FAK1_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MAAAYLDPNLNHTPNSSTKTHLGTGMERSPGAMERVLKVFHYFESNSEPTTWASIIRHGDATDVRGIIQKIVDSHKVKHVACYGFRLSHLRSEEVHWLHVDMGVSSVREKYELAHPPEEWKYELRIRYLPKGFLNQFTEDKPTLNFFYQQVKSDYMLEIADQVDQEIALKLGCLEIRRSYWEMRGNALEKKSNYEVLEKDVGLKRFFPKSLLDSVKAKTLRKLIQQTFRQFANLNREESILKFFEILSPVYRFDKECFKCALGSSWIISVELAIGPEEGISYLTDKGCNPTHLADFTQVQTIQYSNSEDKDRKGMLQLKIAGAPEPLTVTAPSLTIAENMADLIDGYCRLVNGTSQSFIIRPQKEGERALPSIPKLANSEKQGMRTHAVSVSETDDYAEIIDEEDTYTMPSTRDYEIQRERIELGRCIGEGQFGDVHQGIYMSPENPALAVAIKTCKNCTSDSVREKFLQEALTMRQFDHPHIVKLIGVITENPVWIIMELCTLGELRSFLQVRKYSLDLASLILYAYQLSTALAYLESKRFVHRDIAARNVLVSSNDCVKLGDFGLSRYMEDSTYYKASKGKLPIKWMAPESINFRRFTSASDVWMFGVCMWEILMHGVKPFQGVKNNDVIGRIENGERLPMPPNCPPTLYSLMTKCWAYDPSRRPRFTELKAQLSTILEEEKAQQEERMRMESRRQATVSWDSGGSDEAPPKPSRPGYPSPRSSEGFYPSPQHMVQTNHYQVSGYPGSHGITAMAGSIYPGQASLLDQTDSWNHRPQEIAMWQPNVEDSTVLDLRGIGQVLPTHLMEERLIRQQQEMEEDQRWLEKEERFLKPDVRLSRGSIDREDGSLQGPIGNQHIYQPVGKPDPAAPPKKPPRPGAPGHLGSLASLSSPADSYNEGVKLQPQEISPPPTANLDRSNDKVYENVTGLVKAVIEMSSKIQPAPPEEYVPMVKEVGLALRTLLATVDETIPLLPASTHREIEMAQKLLNSDLGELINKMKLAQQYVMTSLQQEYKKQMLTAAHALAVDAKNLLDVIDQARLKMLGQTRPH
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Non-receptor protein-tyrosine kinase that plays an essential role in regulating cell migration, adhesion, spreading, reorganization of the actin cytoskeleton, formation and disassembly of focal adhesions and cell protrusions, cell cycle progression, cell proliferation and apoptosis. Required for early embryonic development and placenta development. Required for embryonic angiogenesis, normal cardiomyocyte migration and proliferation, and normal heart development. Regulates axon growth and neuronal cell migration, axon branching and synapse formation; required for normal development of the nervous system. Plays a role in osteogenesis and differentiation of osteoblasts. Functions in integrin signal transduction, but also in signaling downstream of numerous growth factor receptors, G-protein coupled receptors (GPCR), EPHA2, netrin receptors and LDL receptors. Forms multisubunit signaling complexes with SRC and SRC family members upon activation; this leads to the phosphorylation of additional tyrosine residues, creating binding sites for scaffold proteins, effectors and substrates. Regulates numerous signaling pathways. Promotes activation of phosphatidylinositol 3-kinase and the AKT1 signaling cascade. Promotes activation of MAPK1/ERK2, MAPK3/ERK1 and the MAP kinase signaling cascade. Promotes localized and transient activation of guanine nucleotide exchange factors (GEFs) and GTPase-activating proteins (GAPs), and thereby modulates the activity of Rho family GTPases. Signaling via CAS family members mediates activation of RAC1. Recruits the ubiquitin ligase MDM2 to P53/TP53 in the nucleus, and thereby regulates P53/TP53 activity, P53/TP53 ubiquitination and proteasomal degradation. Phosphorylates SRC; this increases SRC kinase activity. Phosphorylates ACTN1, ARHGEF7, GRB7, RET and WASL. Promotes phosphorylation of PXN and STAT1; most likely PXN and STAT1 are phosphorylated by a SRC family kinase that is recruited to autophosphorylated PTK2/FAK1, rather than by PTK2/FAK1 itself. Promotes phosphorylation of BCAR1; GIT2 and SHC1; this requires both SRC and PTK2/FAK1. Promotes phosphorylation of BMX and PIK3R1. Isoform 6 (FRNK) does not contain a kinase domain and inhibits PTK2/FAK1 phosphorylation and signaling. Its enhanced expression can attenuate the nuclear accumulation of LPXN and limit its ability to enhance serum response factor (SRF)-dependent gene transcription.
Phosphorylated on tyrosine residues upon activation, e.g. upon integrin signaling. Tyr-397 is the major autophosphorylation site, but other kinases can also phosphorylate this residue. Phosphorylation at Tyr-397 promotes interaction with SRC and SRC family members, leading to phosphorylation at Tyr-576, Tyr-577 and at additional tyrosine residues. FGR promotes phosphorylation at Tyr-397 and Tyr-576. FER promotes phosphorylation at Tyr-577, Tyr-861 and Tyr-925, even when cells are not adherent. Tyr-397, Tyr-576 and Ser-722 are phosphorylated only when cells are adherent. Phosphorylation at Tyr-397 is important for interaction with BMX, PIK3R1 and SHC1. Phosphorylation at Tyr-925 is important for interaction with GRB2. Dephosphorylated by PTPN11; PTPN11 is recruited to PTK2 via EPHA2 (tyrosine phosphorylated). Microtubule-induced dephosphorylation at Tyr-397 is crucial for the induction of focal adhesion disassembly; this dephosphorylation could be catalyzed by PTPN11 and regulated by ZFYVE21. Phosphorylation on tyrosine residues is enhanced by NTN1 (By similarity).
Sumoylated; this enhances autophosphorylation.
Cell junction>Focal adhesion. Cell membrane>Peripheral membrane protein>Cytoplasmic side. Cytoplasm>Cell cortex. Cytoplasm>Cytoskeleton. Cytoplasm>Cytoskeleton>Microtubule organizing center>Centrosome. Nucleus.
Note: Constituent of focal adhesions. Detected at microtubules.
Detected in B and T-lymphocytes. Isoform 1 and isoform 6 are detected in lung fibroblasts (at protein level). Ubiquitous.
The Pro-rich regions interact with the SH3 domain of CAS family members, such as BCAR1 and NEDD9, and with the GTPase activating protein ARHGAP26.
The carboxy-terminal region is the site of focal adhesion targeting (FAT) sequence which mediates the localization of FAK1 to focal adhesions.
Belongs to the protein kinase superfamily. Tyr protein kinase family. FAK subfamily.
Research Fields
· Cellular Processes > Cellular community - eukaryotes > Focal adhesion. (View pathway)
· Cellular Processes > Cell motility > Regulation of actin cytoskeleton. (View pathway)
· Environmental Information Processing > Signal transduction > ErbB signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > PI3K-Akt signaling pathway. (View pathway)
· Human Diseases > Drug resistance: Antineoplastic > Endocrine resistance.
· Human Diseases > Infectious diseases: Bacterial > Bacterial invasion of epithelial cells.
· Human Diseases > Infectious diseases: Parasitic > Amoebiasis.
· Human Diseases > Infectious diseases: Viral > Human papillomavirus infection.
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Overview > Transcriptional misregulation in cancer.
· Human Diseases > Cancers: Overview > Proteoglycans in cancer.
· Human Diseases > Cancers: Specific types > Small cell lung cancer. (View pathway)
· Organismal Systems > Immune system > Chemokine signaling pathway. (View pathway)
· Organismal Systems > Development > Axon guidance. (View pathway)
· Organismal Systems > Immune system > Leukocyte transendothelial migration. (View pathway)
References
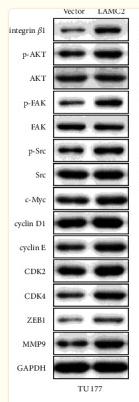
Application: WB Species: Mouse Sample:
Application: IHC Species: Mouse Sample:
Application: WB Species: Human Sample: U87 cells
Application: WB Species: Mouse Sample:
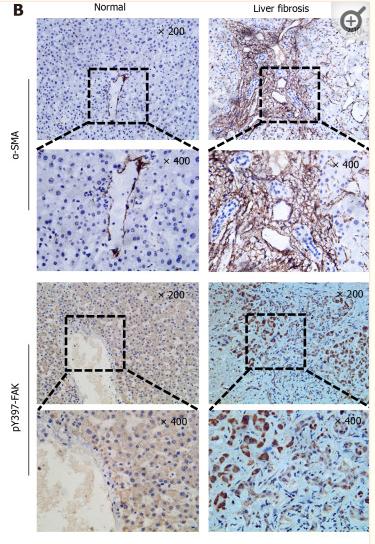
Application: IHC Species: Human Sample: fibrotic liver tissue
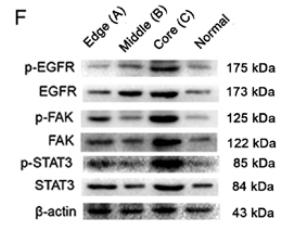
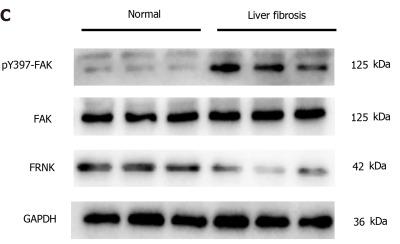
Application: WB Species: Human Sample: fibrotic liver tissue
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.