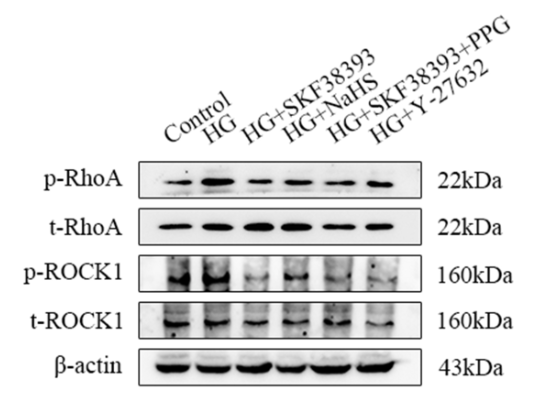
Phospho-RhoA (Ser188) Antibody - #AF3352
| Product: | Phospho-RhoA (Ser188) Antibody |
| Catalog: | AF3352 |
| Description: | Rabbit polyclonal antibody to Phospho-RhoA (Ser188) |
| Application: | WB |
| Cited expt.: | WB |
| Reactivity: | Human, Mouse, Rat, Monkey |
| Prediction: | Pig, Bovine, Sheep, Dog |
| Mol.Wt.: | 22kDa; 22kD(Calculated). |
| Uniprot: | P61586 |
| RRID: | AB_2834767 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF3352, RRID:AB_2834767.
Fold/Unfold
Aplysia ras related homolog 12; ARH12; ARHA; H 12; H12; Oncogene RHO H12; Ras homolog family member A; Ras homolog gene family member A; Rho A; Rho cDNA clone 12; RHO H12; RHO12; RHOA; RHOA_HUMAN; RHOH12; Small GTP binding protein Rho A; Transforming protein Rho A; Transforming protein RhoA;
Immunogens
A synthesized peptide derived from human RhoA around the phosphorylation site of Ser188.
- P61586 RHOA_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MAAIRKKLVIVGDGACGKTCLLIVFSKDQFPEVYVPTVFENYVADIEVDGKQVELALWDTAGQEDYDRLRPLSYPDTDVILMCFSIDSPDSLENIPEKWTPEVKHFCPNVPIILVGNKKDLRNDEHTRRELAKMKQEPVKPEEGRDMANRIGAFGYMECSAKTKDGVREVFEMATRAALQARRGKKKSGCLVL
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Small GTPase which cycles between an active GTP-bound and an inactive GDP-bound state. Mainly associated with cytoskeleton organization, in active state binds to a variety of effector proteins to regulate cellular responses such cytoskeletal dynamics, cell migration and cell cycle. Regulates a signal transduction pathway linking plasma membrane receptors to the assembly of focal adhesions and actin stress fibers. Involved in a microtubule-dependent signal that is required for the myosin contractile ring formation during cell cycle cytokinesis. Plays an essential role in cleavage furrow formation. Required for the apical junction formation of keratinocyte cell-cell adhesion. Essential for the SPATA13-mediated regulation of cell migration and adhesion assembly and disassembly. The MEMO1-RHOA-DIAPH1 signaling pathway plays an important role in ERBB2-dependent stabilization of microtubules at the cell cortex. It controls the localization of APC and CLASP2 to the cell membrane, via the regulation of GSK3B activity. In turn, membrane-bound APC allows the localization of the MACF1 to the cell membrane, which is required for microtubule capture and stabilization. Regulates KCNA2 potassium channel activity by reducing its location at the cell surface in response to CHRM1 activation; promotes KCNA2 endocytosis. May be an activator of PLCE1. In neurons, involved in the inhibiton of the initial spine growth. Upon activation by CaMKII, modulates dendritic spine structural plasticity by relaying CaMKII transient activation to synapse-specific, long-term signaling (By similarity).
(Microbial infection) Serves as a target for the yopT cysteine peptidase from Yersinia pestis, vector of the plague.
(Microbial infection) Substrate for botulinum ADP-ribosyltransferase.
(Microbial infection) Cleaved by yopT protease when the cell is infected by some Yersinia pathogens. This removes the lipid attachment, and leads to its displacement from plasma membrane and to subsequent cytoskeleton cleavage.
(Microbial infection) AMPylation at Tyr-34 and Thr-37 are mediated by bacterial enzymes in case of infection by H.somnus and V.parahaemolyticus, respectively. AMPylation occurs in the effector region and leads to inactivation of the GTPase activity by preventing the interaction with downstream effectors, thereby inhibiting actin assembly in infected cells. It is unclear whether some human enzyme mediates AMPylation; FICD has such ability in vitro but additional experiments remain to be done to confirm results in vivo.
(Microbial infection) Glycosylated at Tyr-34 by Photorhabdus asymbiotica toxin PAU_02230. Mono-O-GlcNAcylation by PAU_02230 inhibits downstream signaling by an impaired interaction with diverse regulator and effector proteins of Rho and leads to actin disassembly.
Phosphorylation by PRKG1 at Ser-188 inactivates RHOA signaling. Phosphorylation by SLK at Ser-188 in response to AGTR2 activation (By similarity).
Ubiquitinated by the BCR(KCTD13) and BCR(TNFAIP1) E3 ubiquitin ligase complexes, leading to its degradation by the proteasome, thereby regulating the actin cytoskeleton and synaptic transmission in neurons.
Cell membrane>Lipid-anchor>Cytoplasmic side. Cytoplasm>Cytoskeleton. Cleavage furrow. Cytoplasm>Cell cortex. Midbody. Cell projection>Lamellipodium. Cell projection>Dendrite.
Note: Localized to cell-cell contacts in calcium-treated keratinocytes (By similarity). Translocates to the equatorial region before furrow formation in a ECT2-dependent manner. Localizes to the equatorial cell cortex (at the site of the presumptive furrow) in early anaphase in an activated form and in a myosin- and actin-independent manner.
The basic-rich region is essential for yopT recognition and cleavage.
Belongs to the small GTPase superfamily. Rho family.
Research Fields
· Cellular Processes > Transport and catabolism > Endocytosis. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Focal adhesion. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Adherens junction. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Tight junction. (View pathway)
· Cellular Processes > Cell motility > Regulation of actin cytoskeleton. (View pathway)
· Environmental Information Processing > Signal transduction > Ras signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Rap1 signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > cGMP-PKG signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > cAMP signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Sphingolipid signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Phospholipase D signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > mTOR signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Wnt signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > TGF-beta signaling pathway. (View pathway)
· Human Diseases > Infectious diseases: Bacterial > Bacterial invasion of epithelial cells.
· Human Diseases > Infectious diseases: Bacterial > Pathogenic Escherichia coli infection.
· Human Diseases > Infectious diseases: Bacterial > Pertussis.
· Human Diseases > Infectious diseases: Bacterial > Tuberculosis.
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Overview > Viral carcinogenesis.
· Human Diseases > Cancers: Overview > Proteoglycans in cancer.
· Human Diseases > Cancers: Overview > MicroRNAs in cancer.
· Human Diseases > Cancers: Specific types > Colorectal cancer. (View pathway)
· Organismal Systems > Immune system > Chemokine signaling pathway. (View pathway)
· Organismal Systems > Circulatory system > Vascular smooth muscle contraction. (View pathway)
· Organismal Systems > Development > Axon guidance. (View pathway)
· Organismal Systems > Immune system > Platelet activation. (View pathway)
· Organismal Systems > Immune system > NOD-like receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > T cell receptor signaling pathway. (View pathway)
· Organismal Systems > Immune system > Leukocyte transendothelial migration. (View pathway)
· Organismal Systems > Nervous system > Neurotrophin signaling pathway. (View pathway)
· Organismal Systems > Endocrine system > Oxytocin signaling pathway.
· Organismal Systems > Digestive system > Pancreatic secretion.
References
Application: WB Species: human Sample:
Application: WB Species: Mouse Sample:
Application: WB Species: Mice Sample: colon tissue
Application: WB Species: Human Sample: HUVECs
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.